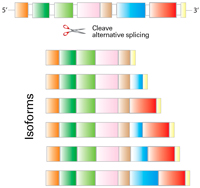
Single-cell transcriptomics (scRNA-seq) has revolutionized the way scientists think about neural organization within the brain, and the Allen Institute for Brain Science has led the way in this field. Researchers at the institute previously used scRNA-seq to analyze single cells at two poles of the mouse neocortex and to define 133 distinct cell types using our SMART-Seq v4 Ultra Low Input Kit for RNA Sequencing (SSv4). In addition, their Patch-seq approach, which compares SSv4 transcriptomic, morphological, and electrophysical analyses for the categorization of brain cells, showed a high correlation between these three parameters. These results validated scRNA-seq as a powerful method for creating brain cell taxonomies. In the most recent work coming out of the institute, Yao and colleagues combine our full-length plate-seq SSv4 kit with a different, yet complementary scRNA-seq approach. The second method was the 3' droplet-based Chromium Single Cell 3' Reagent Kit v2 (10xv2), developed by 10x Genomics. Together, the two techniques unraveled never-before-seen molecular organization within the mouse neocortex and hippocampal formation.
In their recent study published in Cell, the researchers sequenced the transcriptomes of over 1.3 million total individual neurons across the mouse neocortex and hippocampal formation. In all, 1,228,636 cells were analyzed using 10xv2, and 76,381 using SSv4. Clustering data into cell types based on transcriptomic data resulted in an unprecedented 332 clusters using 10xv2 and 324 clusters using SSv4. However, when using a consensus clustering approach to combine the 10xv2 and SSv4 data, the researchers were able to identify 388 unique clusters, 364 of which were neuronal. The additional coverage afforded by combining both data sets allowed the authors to define cell types across the entire neocortex and hippocampal formation without significant gaps and led to the discovery of new cell types in these areas of the brain.
A consensus clustering approach where SSv4 and 10xv2 sequencing data was combined allowed for identification of over 50 additional cell types than clustering from either the SSv4 or 10xv2 sequencing data alone”
| Technique |
SSv4 |
10xv2 |
SSv4 + 10xv2 |
| Total cells sequenced |
76,381 |
1,228,636 |
<1,300,000 |
| Clusters generated |
324 |
332 |
388 |
With this consensus clustering approach, the researchers showed that the “older” hippocampal formation is just as complex, from a cellular composition standpoint, as the more recently evolved neocortex. Additionally, their data demonstrated that cell types across the neocortex and hippocampal formation are organized as a gradient. This means that a given cell in the brain is most like its neighbors, and most unlike cells that are farther away. These findings shed light on how the brain evolved and how it forms during development.
As previous research has clearly demonstrated, both SSv4 and 10xv2 are powerful techniques that promote biological discovery on their own. However, as shown in this study, when these two different yet complementary scRNA-seq approaches are combined, the additional coverage gained provides further biological insight. This allowed researchers at the institute to advance their research goal of gaining a better understanding of the brain through the identification of novel cell types.
And we think that's good science!
References
Yao, Z. et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 184, 3222–3241.e26 (2021).