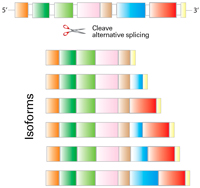
The brain is undoubtedly our most complex organ, with many mysteries still waiting to be unraveled. Founded in 2003, the Allen Institute for Brain Science has embarked on the mission of understanding brain function. Next-generation sequencing (NGS) has been a key tool used by them to help map the brain, and their Single Cell RNASeq Core recently surpassed their milestone of processing over 300,000 samples, all while using Takara Bio's SMART-Seq v4 Ultra Low Input RNA Kit for Sequencing (SSv4).
We sat down with the core's manager, Kimberly Smith, to talk about what it takes to accomplish such a monumental project.

Associate Director, Molecular Biology Kimberly Smith of the Allen Institute
What is the Institute's overall research goal?
The mission of the Allen Institute for Brain Science is to accelerate the understanding of how the brain works in health and disease. To do this, we aim to establish a baseline of what a normal brain is and all its characteristics and components. This year we are planning on completing sequencing the healthy mouse and human brains and transitioning to disease models, such as Alzheimer's.
Tell us about the samples the core processes.
The majority, about 60% of the 300,000 [processed], have been mouse cells or nuclei, most of those being neuronal. We hope to achieve sequencing the entire mouse brain overall. About 35–39% come from human brain tissue. For these, we use exclusively nuclei coming from postmortem brains or surgical tissues. Another 5% of samples are used for Patch-seq, in which live mouse and human cells are examined using electrophysiology rigs, have their nuclei extracted and sequenced for transcriptomics, and are imaged to look at morphology.
How has your approach to tackling such large datasets changed as new technologies emerge?
I've been here since the very beginning, and projects have really morphed to keep up with the emerging technologies ready for production. When we first started, everybody was doing in situ hybridization (ISH) on the mouse brain. For human ISH, we transitioned into microarrays for more targeted analysis. When we first got into RNA sequencing, we weren't working in single cells, but rather in bulk tissue. About 4–5 years ago, we started transitioning into single-cell sequencing.
How many samples a day is that?
We process five 96-wells of amplification plates using the SMART-Seq v4 process. In addition, we process another five plates through the library portion using Illumina Nextera® XT. With controls, per week, we're processing 2,200 samples through both amplification and library.
How has SMART-Seq v4 helped you accomplish your research goal?
We were very happy when [SMART-Seq] v4 came along because it really increased the gene sensitivity for each cell. Our analysis relies on getting high gene content per cell, as we do clustering [analysis] for thousands of cells. The SSv4 kits have been very reliable from lot to lot, giving us the year-to-year consistency critical for this project. As we get shipments in, it just performs with a consistency and reliability that we couldn't get from component-based and in-house-generated approaches such as Smart-seq2, which requires a lot more logistics in terms of monitoring output and making sure it is consistent.
[SMART-Seq] v4 has really been the backbone to what we have been able to accomplish here for its consistency and reliability."
We imagine keeping consistency between users is also a concern. How does that come into play when collecting such large datasets?
We have a set of six trained research associates that take a lot of pride in their work and standardization day to day, but also to ensure that between people we process things in the same detailed fashion. Using manufactured kits like SMART-Seq v4 helps because it helps condense the number of variables (tubes to open, reagents to pipette, etc.) to help maintain consistency.
How many more samples will be processed for this project?
For mouse, we'll probably process about 350,000 samples to finish up missing regions of the brain and developmental timepoints.
For human, we've pretty much wrapped up with current characterization. The next goal is to process samples from many different people. To get those numbers, we will transition to a higher throughput with something like 10x [Genomics]. However, that work wouldn't be nearly as valuable if we didn't have this existing scaffold of 100,000 nuclei already characterized with SMART-Seq v4.
For Patch-seq, we're looking to use the SMART-Seq Single Cell Kit. These cells are very valuable, giving us three data modalities from each cell, so we are interested in maximizing the gene detection, and the single-cell kit lets us extract the most data that we can from these cells.
It's interesting that you started out with full-length and are now moving to higher-throughput technology. Can you comment more on that approach?
They [full-length and high-throughput end capture] are parallel technologies that are complementary."
I think you're missing the whole picture if you do one exclusively over the other. With the SMART-Seq v4, especially for our mouse samples, we can get very high resolution of the tissue that we profile from as few as 1–20 cells, so if we really want to be layer-specific to profile those specialized cells, we can. For higher-throughput technologies, you really need to start with thousands of cells from a large brain region. They both have their advantages and drawbacks, so that is why we use both.
Congratulations to the Allen Institute's sequencing team on this extraordinary achievement!
We are continually inspired to hear about the innovative advancements that researchers like Kimberly and her team are making in their fields with kits like SMART-Seq v4. We are excited to see them continue their efforts to understand the inner workings of the brain.