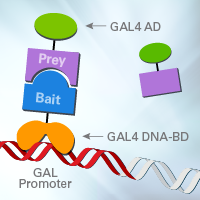
Yeast two-hybrid (Y2H) systems are primarily used for screening a complete library of proteins (prey) for interaction with a specific protein of interest (bait). However, the manufacturing and screening of these libraries has traditionally been both time- and labor-intensive. To address these shortcomings, we've developed our popular, ready-to-go Mate & Plate libraries that make Y2H screens simple: co-culture your library strain with your bait strain, plate on the appropriate selective minimal media, and you're done.
Tech Note
Make your own library for yeast two-hybrid screening
- Library construction directly in yeast using SMART technology
- No laborious cloning or library amplification steps
- Enough material for hundreds of yeast two-hybrid screens
Featured products: ♦ Mate & Plate libraries ♦ Make Your Own "Mate & Plate" Library System ♦ Advantage 2 Polymerase Mix ♦ Matchmaker AD LD-Insert Screening Amplimer Set
Introduction
Do-it-yourself Y2H libraries
If our selection of ready-made libraries doesn't suit your needs, we offer the Make Your Own "Mate & Plate" Library System. This kit provides the necessary materials and a simple, highly efficient method to make enough Mate & Plate library for hundreds of Y2H screens, all in less than a week.
Library creation occurs directly in our library Y187 Yeast Strain, utilizing the highly efficient homologous recombination machinery of S. cerevisiae (Figure 1, Panel A). This system uses our proprietary SMART cDNA synthesis technology (Figure 1, Panel B), allowing you to construct cDNA libraries from any tissue source with as little as 100 ng of total input RNA. This eliminates the need for the labor-intensive library cloning, amplification, and harvesting in E. coli required in traditional library construction methods.

Figure 1. Library generation using in vivo recombination and SMART technology in S. cerevisiae. SMART cDNA synthesis generates cDNAs with homologous ends (Panel A), enabling the creation of Mate & Plate libraries via recombination between your cDNA and the Matchmaker prey vector pGADT7-Rec (Panel B). Following this, colonies are pooled, mixed, and aliquoted into multiple vials.
High-efficiency cloning with yeast
The highly efficient homologous recombination pathways of S. cerevisiae yeast have been well-documented (Pâques and Haber 1999; Sung et. al. 2000). Their efficiency has been exploited for decades to enable E. coli-free cloning of plasmids through a process known as gap-repair (Ma et al. 1987). We've taken this one step further by providing a way for you to perform E. coli-free cloning of an entire library directly in yeast. The entire process consists of four simple steps:
Step 1: first-strand synthesis using the included SMART oligo to generate cDNA homologous at both ends to pGADT7-Rec
Step 2: second-strand PCR synthesis to generate 2–5 µg of library cDNA
Step 3: cotransformation and recombination in the Y187 Yeast Strain
Step 4: harvesting, mixing, and aliquoting of colonies into 1-ml single-use vials, providing enough material for hundreds of Y2H screens
Enrichment for longer cDNA clones
The cDNA size range for inserts cloned with this protocol is 0.3–6 kb (Figure 2). As the SMART oligo sequence is required for homology to the prey vector, prematurely terminated reverse transcripts are selected against because they cannot be amplified using the second-strand synthesis primers. Our Make Your Own "Mate & Plate" Library System also contains CHROMA SPIN gel filtration columns to size-select for larger (>400 bp) cDNAs (Figure 2) and deliver complex, representative libraries (Figure 3).

Figure 2. High-quality cDNA and complex, representative libraries generated with SMART cDNA synthesis. Oligo dT-primed cDNA was generated using the Make Your Own "Mate & Plate" Library System with (positive control) or without (negative control) 1 µg of human placenta polyA RNA. LD PCR was performed using the Advantage 2 Polymerase Mix (with duplicate samples). One set of products was purified and size-selected using CHROMA SPIN+TE-400 columns, and 10 µl of each sample was run on a 1% agarose gel. Lanes M: 1-kb DNA ladder size marker. Lanes 1 and 3 represent unpurified and purified negative controls, respectively. Lanes 2 and 4 represent unpurified and purified positive controls, with Lane 4 showing a reduced abundance of cDNA species of <400 bp.

Figure 3. Mate & Plate libraries display broad insert representation. A human bone marrow library was made using the Make Your Own "Mate & Plate" Library System. Inserts from 15 randomly picked colonies were analyzed by yeast colony PCR using the Advantage 2 Polymerase Mix and the Matchmaker AD LD-Insert Screening Amplimer Set. As seen in Lanes 1–15, each colony contained an insert of a different size. Lane M: 1-kb DNA ladder size marker.
References
Ma, H., Kunes, S., Schatz, P. J. & Botstein, D. Plasmid construction by homologous recombination in yeast. Gene 58, 201–16 (1987).
Pâques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 63, 349–404 (1999).
Sung, P., Trujillo, K. M. & Van Komen, S. Recombination factors of Saccharomyces cerevisiae. Mutat. Res. 451, 257–75 (2000).
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.