Cogent™ NGS Analysis Pipeline
Version 3.2
Overview
Cogent NGS Analysis Pipeline (also referred to as CogentAP) is bioinformatic software for analyzing single-cell RNA-seq, bulk RNA-seq, and single-cell DNA-seq data. Functionalities include gene or transcript expression counting, in silico ribodepletion, gene fusion detection, immune profiling analysis, and more. Please refer to the table in the bioinformatics resource portal for the full list of validated Takara Bio reagent kits supported for use with the pipeline. CogentAP inputs raw sequencing data and outputs an HTML report and other files, such as a gene matrix, to continue further analysis. Output also includes R data objects, which may be used as input for Cogent NGS Discovery Software.

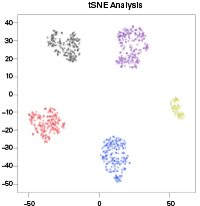
Example output available in the CogentAP HTML report.
Install the software to access the following features:
- Command-line interface program
- Mini-dataset files (WellList.txt and FASTQ files)—used to test the CogentAP functionality
- Sample output reports and stats—used to compare to the analysis results of the mini-dataset test
More Information
Supported operating systems
CogentAP can be installed on the following versions of Linux:
- CentOS 8 or higher
- Red Hat 8 or higher
- Ubuntu 18.4 or higher
Hardware requirements
For analyzing Illumina NextSeq® High-Output data:
- 24-core CPU
- 64 GB+ RAM
- 1 TB available hard drive space*
Analyzing MiniSeq® or MiSeq® data may be possible on servers with less computational power than the specifications above. Analysis of HiSeq® or NovaSeq™ data would require better hardware than is listed.
*Data generated using the Shasta Total RNA-Seq Kit or Shasta Whole-Genome Amplification Kit requires at least six times the size of the input FASTQ files of free hard drive space for analyses with default parameters, and at least ten times the size of the input FASTQ files of free hard drive space for analyses including ribodepletion, immune profiling, and fusion analyses.
Additional third-party software dependencies*
- Conda
- bcl2fastq or BCL Convert for conversation of sequencer FASTQ output files
*Not included with the software; must be downloaded and installed separately.
Supported applications
See a list of supported applications.
Additional product information
More information is available in the Cogent NGS Analysis Pipeline User Manual or Quick Start Guide.
Cogent NGS Analysis Pipeline version history
| Version number | Release date |
Notes |
|---|---|---|
| CogentAP v3.2 | 2025-08-01 |
|
| CogentAP v3.1.27 | 2025-04-23 |
|
| CogentAP v3.1.23 | 2025-04-02 |
|
| CogentAP v3.1 | 2025-01-15 |
|
|
CogentAP 2.0.1 |
2023-05-26 |
|
| CogentAP 2.0 | 2022-08-17 |
|
| CogentAP 1.5.1 | 2022-03-25 |
|
| CogentAP 1.5 | 2021-12-03 |
|
| CogentAP 1.0 | 2020-09-11 |
|
| mappa Analyzer 1.0 | 2019-06-07 |
|

More cells. More biomarkers. More discoveries.
Want to go from sample prep through data analysis with less hands-on time and stress? Meet the Shasta Single Cell System, which helps researchers find novel biomarkers. Learn how our high-throughput Shasta technologies can inspire the next breakthrough.
Get the details
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.