SMARTer Human TCR a/b Profiling Kit v2—flexible kit with ultimate sensitivity for understanding the TCR repertoire

NOTE: We recommend the reconfigured versions of the kits below, SMART-Seq Human TCR (with UMIs) (Cat. # 634780, 634781 & 634779).
SMARTer Human TCR a/b Profiling Kit v2 (Cat. # 634478, 634479) will be phased out soon. Contact your local sales representative for more information.
The SMARTer Human TCR a/b Profiling Kit v2 (TCRv2) is powered by robust chemistry that provides unparalleled sensitivity and reproducibility. The kit leverages SMART (Switching Mechanism at 5' end of RNA Template) full-length cDNA synthesis technology and pairs NGS with a 5'-RACE approach to capture the complete V(D)J variable regions of TRA and TRB genes.
NOTE: We recommend the reconfigured versions of the kits below, SMART-Seq Human TCR (with UMIs) (Cat. # 634780, 634781 & 634779).
SMARTer Human TCR a/b Profiling Kit v2 (Cat. # 634478, 634479) will be phased out soon. Contact your local sales representative for more information.
The SMARTer Human TCR a/b Profiling Kit v2 (TCRv2) is powered by robust chemistry that provides unparalleled sensitivity and reproducibility. The kit leverages SMART (Switching Mechanism at 5' end of RNA Template) full-length cDNA synthesis technology and pairs NGS with a 5'-RACE approach to capture the complete V(D)J variable regions of TRA and TRB genes.
The kit is designed to work with a range of RNA input amounts (RIN ≥8; depending on the sample type) and has been shown to yield high-quality sequencing libraries from as little as 10 ng to 1 µg of total RNA obtained from peripheral blood leukocytes, 20 ng to 200 ng of total RNA obtained from whole blood, 1 ng to 100 ng of total RNA obtained from T cells, or from 1,000 to 10,000 purified, whole T cells. Libraries can be generated to obtain both alpha and beta chain diversity information. This latest profiling kit also includes unique molecular identifiers (UMI), making it possible to remove reads derived from PCR duplicates and sequencing errors, thus ensuring more accurate and reliable results. In addition to the reagents, the TCRv2 kit includes a dual index (UDI) plate to generate Illumina-compatible sequencing libraries.
Benefits of this kit include:
- Use of UMIs to correct for PCR and sequencing errors
- Addition of UDIs, allowing for greater confidence in sequencing on a patterned flow cell (such as the NovaSeq™ system) and the ability to pool a greater number of samples
- Flexibility to sequence on any Illumina instrument
- Full-length V(D)J analysis on the MiSeq™ system
Overview
- Compatible with a wide range of sample inputs: total RNA (10 ng to 1 µg from peripheral blood leukocytes, 20 ng to 200 ng of total RNA obtained from whole blood, or 1 ng to 100 ng from T cells) or purified, whole T cells (1,000 to 10,000 cells)
- Simple PCR amplification: a single primer pair for each TCR (alpha or beta) subunit per reaction
- UMI-based correction: removal of reads derived from PCR duplicates and sequencing errors
- Sensitive and specific clonotype detection: optimized cDNA library generation
- UDI implementation: increased multiplexing and confidence for sequencing on high-throughput sequencers
- Illumina-ready sequencing libraries: Illumina-compatible index sequences are incorporated for multiplexing up to 192 libraries in a single run
- Flexible sequencing options: either full-length V(D)J information on the MiSeq™ system or CDR3 information on all Illumina platforms

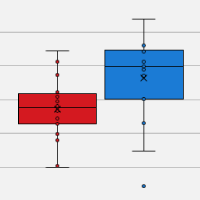
Figure 1. Sensitive and reproducible clonotype detection from a broad range of RNA amounts. TRA and TRB libraries were generated from 1, 10, and 100 ng of human CD3+ T-cell total RNA and 1,000 and 10,000 CD3+ T cells. The sequence reads were processed by the Cogent NGS Immune Profiler Software.
| % Jurkat RNA spiked in to 100 ng of PBMC RNA | Total read count (TRA/TRB) | Without UMI collapse | With UMI collapse | ||||
|---|---|---|---|---|---|---|---|
| # of TRB raw reads | # of reads for TRBV12‑3-TRBJ1‑2 | Detected percentage of Jurkat reads | # of detected UMIs | # of UMIs for TRBV12‑3-TRBJ1‑2 | Detected percentage of Jurkat UMIs | ||
| 10.0% | 2,500,000 | 1,565,005 | 397,179 | 25.0% | 281,280 | 62,629 | 22.0% |
| 1.0% | 2,500,000 | 1,422,102 | 47,160 | 3.3% | 219,776 | 6,426 | 2.9% |
| 0.1% | 2,500,000 | 1,366,127 | 5,412 | 0.4% | 189,580 | 631 | 0.33% |
| 0.01% | 2,500,000 | 1,218,025 | 521 | 0.043% | 196,615 | 74 | 0.038% |
| 0.001% | 2,500,000 | 1,331,465 | 909 | 0.068% | 197,870 | 6 | 0.003% |
| 0.0001% | 2,500,000 | 1,409,199 | - | 0% | 124,149 | - | 0% |
| 0% | 2,500,000 | 1,222,245 | - | 0% | 197,933 | - | 0% |
Table 1. Assessing the sensitivity and reproducibility of the SMARTer approach. Spike-in analysis was performed in replicate on PBMC RNA samples spiked at varying concentrations (10%, 1%, 0.1%, 0.01%, 0.001%, and 0.0001%) with RNA obtained from a homogeneous population of leukemic Jurkat T cells (containing TRBV12-3-TRBJ1-2 clonotypes). TRB CDR3 regions were amplified from 100 ng of total RNA using the TCRv2 kit and sequenced. Reads of 2 x 150 bp were obtained on an Illumina NextSeq® system. The sequencing reads were downsampled to 2.5 M reads. Read results for spike-in concentrations identified as the reliable concentration limit for each criterion (without and with UMI collapse) have data highlighted in gray. Without UMI collapse, PCR duplicates of TRBV12-3 were observed in 0.0010% of the raw reads.

Figure 2. Superior sensitivity and reproducibility with TCRv2. We split 5M PBMC cells from two different healthy donors for RNA and gDNA extraction. 1.6 µg of gDNA was used for library preparation according to the manufacturer's instructions (15% of the total amount of extracted gDNA). 100 ng of RNA was used for library preparation (2% of the total amount of extracted RNA). Clonotype numbers for TCRa/b libraries are shown from each company (NT: not tested). In the comparison, TCRv2 generated an average of 48.7K and 163K clonotypes for TRA and TRB, respectively, representing a 290% increase against Company Q's RNA-based approach and a 145% increase against Company A's gDNA-based approach. Importantly, the RNA methods used only 2% of the total RNA from the 5M PBMCs.
More Information
Applications
Human TCR repertoire analysis (TCRα and TCRβ subunits)
Additional product information
Please see the product's Certificate of Analysis for information about storage conditions, product components, and technical specifications. Please see the Kit Components List to determine kit components. Certificates of Analysis and Kit Components Lists are located under the Documents tab.
Product line overview
Browse our broad NGS portfolio, which provides unmatched sensitivity for all of your demanding sequencing applications.
RNA-seq learning center
Learn more about our RNA-seq technology and applications.
Tips for BCR/TCR profiling
Get started with NGS-based immune profiling of B- and/or T-cell repertoires
Technical notes
Technical notes for a wide variety of NGS applications.
Technology and application overviews
Overviews of our NGS technologies and applications.
FAQs and tips
Helpful information for successful NGS experiments.
DNA-seq protocols
Currently featuring protocols for integrating ThruPLEX kits with leading target enrichment systems.
Bioinformatics resources
Browse our library of bioinformatics software resources, including kit and reagent compatibility, user manuals, software notices, troubleshooting tips, and videos.
Webinars
Prerecorded webinars covering a variety of NGS applications.
Posters
Posters featuring Takara Bio NGS methods and data.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.