Choosing the right tool for designing guide RNAs
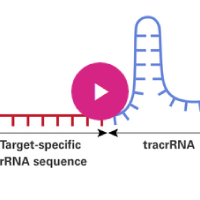
The first step of CRISPR/Cas9 gene editing is designing a single guide RNA (sgRNA) to target your gene of interest. Because sgRNAs are solely responsible for recruiting Cas9 to specific genomic loci, optimal sgRNA design is critical for successful gene editing experiments. There are many web-based tools available for sgRNA design, each of which has different features and advantages. The information provided here will help you choose the best tool for your specific research objective.
Several web-based tools available
Web-based sgRNA design tools typically require that users input a DNA sequence, genomic location, or gene name for each gene of interest, and indicate a species. An algorithm specific to each tool outputs a list of candidate guide sequences with corresponding predicted off-target sites for each input (Wu et al. 2014). Most tools aim to provide guide sequences that minimize the likelihood of off-target effects, but the methods they employ vary. For example, Chop Chop uses empirical data from multiple recent publications (e.g. Doench et al. 2016) to calculate efficiency scores. Alternatively, CasFinder (Aach et al. 2014) and E-CRISP (Heigwer et al. 2014) incorporate specific user-defined penalties based on the number and position of mismatches relative to the guide sequence in order to rank the potential for off-target effects.
Tools for specific applications
Some sgRNA design tools have been developed for specific applications. CRISPR-ERA (Liu et al. 2015) is the only currently available tool that designs sgRNAs specifically for gene repression or activation, while FlyCRISPR (Gratz et al. 2013) focuses on applications in fly, beetle, and worm species, including the popular model organisms Drosophila melanogaster and Caenorhabditis elegans. Presently, the design tool featured on the Benchling website is the only one that can generate candidate sgRNAs that are compatible with alternative nucleases such as Staphylococcus aureus Cas9 (Ran et al. 2015) and Cpf1 (Zetsche et al. 2015). Given the uniqueness of each tool, we recommend that you use multiple approaches during the sgRNA design process and choose guide sequences that are consistently predicted to perform well.
A selection of freely available tools
The table below provides a list of web-based tools for sgRNA design. For simplicity, we have only included those that are free and do not require a subscription. For each tool, we have indicated whether there is a convenient graphical user interface or if the user has to download a script. If you would prefer to design sgRNAs manually, please visit Choosing a target sequence for CRISPR/Cas9 gene editing to learn more.
| Noteworthy features of web-based sgRNA design tools | |||||
|---|---|---|---|---|---|
| Name | Graphical user interface | Available species | Input | Output | Ranked list |
| CHOPCHOP Citation » |
Yes | 23 | DNA sequence, gene name, genomic location | Candidate guide sequences and off-target loci | Yes |
| CasFinder Citation » |
No (Perl script) | User input | DNA sequence | Candidate guide sequences and off-target loci | Yes |
| Cas-OFFFinder Citation » |
Yes | 11 | Guide sequence | Off-target loci for guide sequences | No |
| FlyCRISPR Citation » |
Yes | 18 | DNA sequence | Candidate guide sequences and off-target loci | No |
| E-CRISP Citation » |
Yes | 31 | DNA sequence or gene name | Candidate guide sequences and off-target loci | Yes |
| CRISPR-ERA Citation » |
Yes | 9 | DNA sequence, gene name, or TSS location | Candidate guide sequences and distances to TSS | Yes |
| Benchling | Yes | 5 | DNA sequence or gene name | Candidate guide sequences and off-target loci | Yes |
References
Aach, et al. Flexible algorithm for identifying specific Cas9 targets in genomes. BioRxiv, Cold Spring Harbor Labs. doi: http://dx.doi.org/10.1101/005074 (2014).
Bae, et al. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics. 30(10):1473–1475 (2014).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotech 34, 184–191 (2016).
Gratz, et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics. 196(4):961–971 (2014).
Heigwer, et al. E-CRISP: fast CRISPR target site identification. Nat Methods. 11(2):122–123 (2014).
Montague, et al. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42(W1):W401–W407 (2014).
Liu, et al. CRISPR-ERA: a comprehensive design tool for CRISPR-mediated gene editing, repression and activation. Bioinformatics. 31(22):3676–3678 (2015).
Ran, et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature. 520(7546):186–191 (2015).
Wu, et al. Target specificity of the CRISPR-Cas9 system. Quant Biol. 2(2):59–70 (2015).
Zetsche, et al. Cpf1 is a single RNA-guided endonuclease of a Class 2 CRISPR-Cas System. Cell. 163(3):759–771 (2015).
CRISPR/Cas9 information
Choosing sgRNA design tools
Browse a collection of sgRNA design tools for Cas9-based genome editing experiments.
Choosing a target sequence for CRISPR/Cas9 gene editing
Learn how to design sgRNA sequences for successful gene editing.
The CRISPR/Cas9 system for targeted genome editing
Overview of CRISPR/Cas9 system for genome editing.
CRISPR/Cas9 genome editing tools
An overview of tools available for each step in a successful genome editing workflow.
Gene editing technical notes
Delivery of Cas9 and sgRNA to mammalian cells using a variety of innovative tools.
SNP engineering application note
Learn about a simple assay for sensitive detection of single-nucleotide substitutions in bulk-edited or clonal cell populations.
CRISPR/Cas9 gesicles overview
Learn about Guide-it CRISPR/Cas9 Gesicle Production System components and workflow.
CRISPR library screening webinar
Watch this webinar to learn how you can perform genome-wide lentiviral sgRNA screens easily.
Choosing an HDR template format
Watch a webinar on how to choose the right HDR template for knockin experiments.
Guide-it SNP Screening Kit FAQs
Get answers to frequently asked questions and view a video explaining the enzymatic assay.
Which are the best web and mobile apps to advance your research? We have your list!
Check out these convenient research tools for PCR, cloning, CRISPR, virus prep, and more.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.