How to design sgRNA sequences
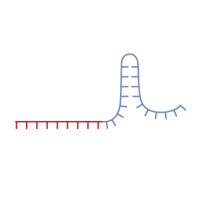
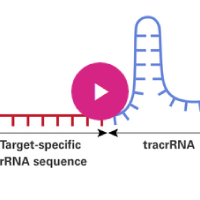
CRISPR/Cas9 gene targeting requires a custom single guide RNA (sgRNA) that contains a targeting sequence (crRNA sequence) and a Cas9 nuclease-recruiting sequence (tracrRNA). The crRNA region (shown in red below) is a 20-nucleotide sequence that is homologous to a region in your gene of interest and will direct Cas9 nuclease activity.
Watch the video to learn more...
Use the following guidelines to select a genomic DNA region that corresponds to the crRNA sequence of the sgRNA:
The 3' end of the DNA target sequence must have a proto-spacer adjacent motif (PAM) sequence (5'-NGG-3'). The 20 nucleotides upstream of the PAM sequence will be your targeting sequence (crRNA), and Cas9 nuclease will cleave approximately three bases upstream of the PAM.
- The PAM sequence itself is absolutely required for cleavage, but it is NOT part of the sgRNA sequence and therefore should not be included in the sgRNA.
- The target sequence can be on either DNA strand.

There are several online tools that detect PAM sequences and list possible crRNA sequences within a specific DNA region. These algorithms also predict off-target effects elsewhere in the genome, allowing you to choose the most specific crRNA for your research application. Please visit our page about sgRNA design tools to learn more.
Through our own experience, we have identified additional tips for designing sgRNAs. We have found that the best sgRNAs for several tested genes have a G at position 1 and an A or T at position 17.
Tips for designing sgRNAs
Synthesize your sgRNA with the Guide-it sgRNA In Vitro Transcription Kit. With this kit, PCR is used to generate a template DNA containing the sgRNA-encoding sequence (constituted by your crRNA and a tracRNA) under the control of a T7 promoter. Then, in vitro transcription using the PCR product template is used to produce sgRNAs that can be purified for efficiency testing and/or cell transduction.
Producing sgRNAs
Even if you pick a target sequence that fulfills all of the described requirements, sgRNA specificity and activity is unpredictable. Therefore, it is often recommended that multiple, different sgRNAs are designed to target a gene of interest. Our Guide-it sgRNA Screening Kit provides a simple and robust in vitro method for assessing sgRNA efficiency before cell transduction, allowing you to identify the most effective CRISPR.
Using the most effective sgRNA
Learn more about Guide-it products for genome editing »
CRISPR/Cas9 information
Choosing sgRNA design tools
Browse a collection of sgRNA design tools for Cas9-based genome editing experiments.
Choosing a target sequence for CRISPR/Cas9 gene editing
Learn how to design sgRNA sequences for successful gene editing.
The CRISPR/Cas9 system for targeted genome editing
Overview of CRISPR/Cas9 system for genome editing.
CRISPR/Cas9 genome editing tools
An overview of tools available for each step in a successful genome editing workflow.
Gene editing technical notes
Delivery of Cas9 and sgRNA to mammalian cells using a variety of innovative tools.
SNP engineering application note
Learn about a simple assay for sensitive detection of single-nucleotide substitutions in bulk-edited or clonal cell populations.
CRISPR/Cas9 gesicles overview
Learn about Guide-it CRISPR/Cas9 Gesicle Production System components and workflow.
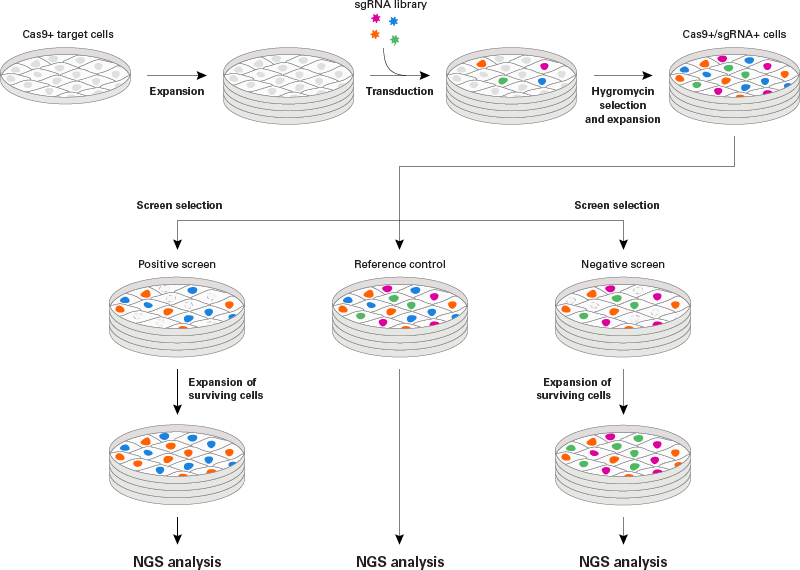
CRISPR library screening webinar
Watch this webinar to learn how you can perform genome-wide lentiviral sgRNA screens easily.
Choosing an HDR template format
Watch a webinar on how to choose the right HDR template for knockin experiments.
Guide-it SNP Screening Kit FAQs
Get answers to frequently asked questions and view a video explaining the enzymatic assay.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.