Technical notes
As the use of single-cell NGS advances, the parallel processing of single cells at high throughputs will be critical. The ICELL8 Single-Cell System and ICELL8 cx Single-Cell System automate single-cell isolation, selection, and processing, allowing for the scalability and simplification of single-cell library preparation. Powerful benefits of using this technology, which increase the versatility of the platforms, include the ability to visualize cells in the chip nanowells and the flexibility to dispense into them multiple times. Furthermore, adaptations of our SMARTer NGS chemistries for RNA library preparation on the ICELL8 systems enable fast, reliable, high-throughput cDNA synthesis for downstream NGS applications. This pairing provides results with high similarity to those of manually generated libraries while increasing reproducibility and scalability. While the systems come validated with various applications, they are open platforms that allow the user to develop their own applications.
Enhancing biomarker discovery with the SMART-Seq Pro kit and ICELL8 cx Single-Cell System
Boost the speed and power of your research with the SMART-Seq Pro application kit, which improves your ability to detect important biomarkers and scale your current full-length experiments.
Target enrichment with the ICELL8 full-length workflow for superior fusion detection
Learn more about improving rare isoform and fusion detection based on the SMART-Seq ICELL8 cx application.
Full-length scRNA-seq for better detection of gene fusions, SNPs, and alternative splicing
Learn how the ICELL8 cx SMART-Seq protocol generates full-length scRNA-seq libraries that enable deeper analyses—such as detection of gene fusions, SNPs, and alternative splicing—than 3' DE on droplet-based systems.
Full-length, single-cell RNA-seq with the ICELL8 Single-Cell System
Learn about automated single-cell RNA-seq library generation for full-length transcriptome analysis from hundreds of single cells in parallel.
User-generated protocol: ATAC-Seq on the ICELL8 Single-Cell System
Sign up to access the user-generated protocol for performing ATAC-Seq on the ICELL8 single-cell platforms (developed by the Greenleaf lab at Stanford University in consultation with TBUSA).
A high-throughput method for single-cell ATAC-seq on the ICELL8 Single-Cell System
Learn more about scATAC-seq on the ICELL8 Single-Cell System.
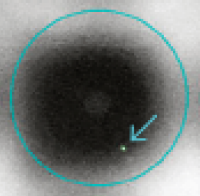
Single-cell identification with CellSelect Software
See how the ICELL8 platform's Imaging System and CellSelect Software allow for high-fidelity identification of individual cells.
Single-cell analysis elucidates cardiomyocyte differentiation from human induced pluripotent stem cells
Learn how researchers used the ICELL8 Single-Cell System to develop a method for transcriptomically characterizing the differentiation of human iPSCs into cardiomyocytes.
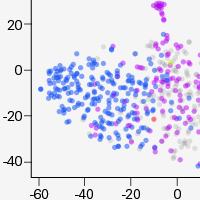
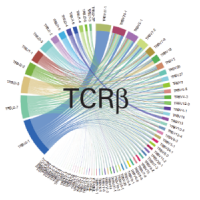
High-throughput TCR profiling with 5'-end differential analysis of single cells
View data on high-throughput whole transcriptome and TCR profiling data from the same cell using the ICELL8 systems.
High-throughput single-cell T-cell receptor profiling with SMART technology
View single-cell TCR profiling data generated with SMART technology and the ICELL8 Single-Cell System.
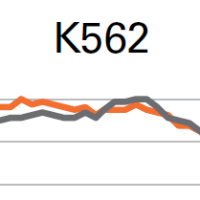
What's inside automated single-cell RNA-seq platforms?
Learn how the many differences between popular scRNA-seq platforms impact your data. Here, we discuss sensitivity, read efficiency, and gene diversity (3' DE expression) data generated by the ABRF Genomics Research Group.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.