Introduction to the CRISPR/Cas9 system
A powerful method for engineering your gene of interest
Although recently developed programmable editing tools, such as zinc finger nucleases and transcription activator-like effector nucleases, have significantly improved the capacity for precise genome modification, these techniques have limitations. CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 technology represents a significant improvement over these other next-generation genome editing tools, reaching a new level of targeting, efficiency, and ease of use. The CRISPR/Cas9 system allows for site-specific genomic targeting in virtually any organism.
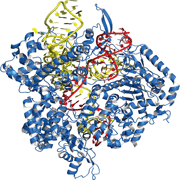
The type II CRISPR/Cas system is a prokaryotic adaptive immune response system that uses noncoding RNAs to guide the Cas9 nuclease to induce site-specific DNA cleavage. This DNA damage is repaired by cellular DNA repair mechanisms, either via the non-homologous end joining DNA repair pathway (NHEJ) or the homology-directed repair (HDR) pathway.
The CRISPR/Cas9 system has been harnessed to create a simple, RNA-programmable method to mediate genome editing in mammalian cells, and can be used to generate gene knockouts (via insertion/deletion) or knockins (via HDR). To create gene disruptions (Figure 1), a single guide RNA (sgRNA) is generated to direct the Cas9 nuclease to a specific genomic location. Cas9-induced double strand breaks are repaired via the NHEJ DNA repair pathway. The repair is error-prone, and thus insertions and deletions (INDELs) may be introduced that can disrupt gene function.

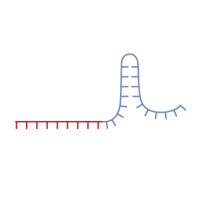
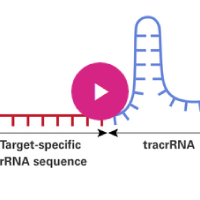
The principle of CRISPR/Cas9-mediated gene disruption. A single guide RNA (sgRNA), consisting of a crRNA sequence that is specific to the DNA target, and a tracrRNA sequence that interacts with the Cas9 protein (1), binds to a recombinant form of Cas9 protein that has DNA endonuclease activity (2). The resulting complex will cause target-specific double-stranded DNA cleavage (3). The cleavage site will be repaired by the nonhomologous end joining (NHEJ) DNA repair pathway, an error-prone process that may result in insertions/deletions (INDELs) that may disrupt gene function (4).
CRISPR/Cas9 technology has revolutionized genome editing, allowing a previously unattainable level of genomic targeting, efficiency, and simplicity. Guide-it products further improve the usability of the CRISPR/Cas9 system by providing a streamlined method for:
CRISPR/Cas9 information
Choosing sgRNA design tools
Browse a collection of sgRNA design tools for Cas9-based genome editing experiments.
Choosing a target sequence for CRISPR/Cas9 gene editing
Learn how to design sgRNA sequences for successful gene editing.
The CRISPR/Cas9 system for targeted genome editing
Overview of CRISPR/Cas9 system for genome editing.
CRISPR/Cas9 genome editing tools
An overview of tools available for each step in a successful genome editing workflow.
Gene editing technical notes
Delivery of Cas9 and sgRNA to mammalian cells using a variety of innovative tools.
SNP engineering application note
Learn about a simple assay for sensitive detection of single-nucleotide substitutions in bulk-edited or clonal cell populations.
CRISPR/Cas9 gesicles overview
Learn about Guide-it CRISPR/Cas9 Gesicle Production System components and workflow.
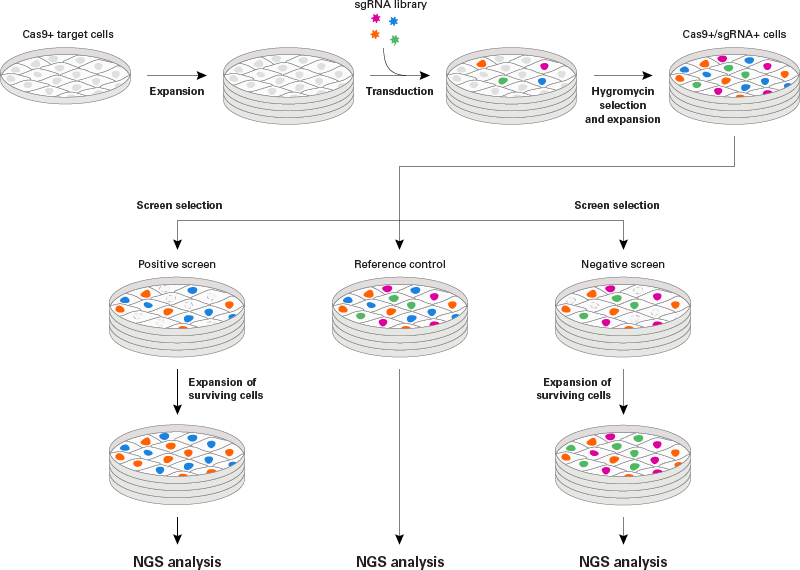
CRISPR library screening webinar
Watch this webinar to learn how you can perform genome-wide lentiviral sgRNA screens easily.
Choosing an HDR template format
Watch a webinar on how to choose the right HDR template for knockin experiments.
Guide-it SNP Screening Kit FAQs
Get answers to frequently asked questions and view a video explaining the enzymatic assay.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.