CRISPR/Cas9 knockins
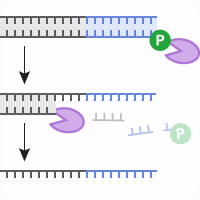
In addition to creating indels or knockouts, scientists can encourage a precise form of repair (homology-directed repair; HDR) by providing a DNA sequence that the cell can use as a repair template to insert (knock in) a matching DNA sequence into the break. Example applications include modification of a promoter sequence or gene, insertion of an exogenous reporter (e.g., a fluorescent protein), or creation of a clincally relevant SNP for a disease model. It should be noted that knockin efficiency is usually lower than knockout efficiency. Moreover, knockin at one copy of the gene/sequence is usually accompanied by a knockout/indel at the second copy; both copies are cut by the Cas9/sgRNA complex but one is likely repaired imprecisely using the non-homologous end joining (NHEJ) pathway, creating an indel.
The links below provide general information, tech notes, and tools used to create or study gene knockins. Although many template formats (plasmids, PCR products, viruses, transposons, etc.) can be used for HDR, the use of single-stranded DNA (ssDNA) templates is increasing rapidly due to the growing understanding that ssDNA integration is far more specific and less toxic than double-stranded DNA (dsDNA) templates.
Choose an HDR template format
Watch a webinar on how to choose the right HDR template for knockin experiments.
Homology-directed repair FAQs
Maximize your likelihood for success by reading these FAQs compiled by our R&D team.
Gene editing of T cells and HSCs
Follow these protocols developed by our R&D team for easy CRISPR/Cas-based gene editing of immune cells.
Site-specific gene knockins using long ssDNA
Gene knockins with long ssDNA sequences produced using Guide-it Long ssDNA Production System v2.
CRISPR/Cas9-mediated knockins in iPS cells
Data demonstrating efficient genome editing of iPS cells using HDR templates generated with the Guide-it Long ssDNA Production System.
CRISPR takes a giant step towards the clinic
Learn about the development of a powerful new method for reprogramming T cells.
Mouse CRISPR knockin protocol
Access a customer-developed protocol for precise genome editing in mouse embryos.
Electroporation-grade Cas9 for editing in diverse cell types
Our Cas9 performs highly efficient gene editing, including in iPS and hematopoietic stem cells.
Screening for effective guide RNAs
Before delivering sgRNA to your cells, use a novel in vitro assay to get accurate predictions of sgRNA cleavage efficiency.
Tag an endogenous gene with AcGFP1 in iPS cells
Workflow for tagging endogenous genes to generate reporter lines.
Tag an endogenous gene with a myc tag in iPS cells
Workflow for tagging endogenous genes with an epitope tag.
Efficient SNP engineering
Learn about a simple assay for sensitive detection of single-nucleotide substitutions in bulk-edited or clonal cell populations.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.