American Society of Gene and Cell Therapy
Radically transform your gene editing workflow
Simplified tools with elegant efficiency to fuel your next breakthrough
![]()
The ASGCT Annual Meeting 2025 happened May 13–17 in New Orleans, LA, and brought professionals together to learn about important issues and advancements in gene and cell therapy. Explore our posters to learn about our comprehensive solutions across viral- and protein-based gene delivery that support research developing the next generation of cell and gene therapies.
Not affiliated with or endorsed by ASGCT.
Featured talk
Streamlined ex vivo engineering of human T cells: A single-step approach to activation and lentiviral transduction

In this talk, Tom Quinn presented an innovative approach to enhance the ex vivo engineering of human T cells by integrating cell activation and lentiviral transduction into a single, efficient step, reducing time and resource requirements while maintaining high performance. The Lenti-X T-Cell Transduction Sponge provides a faster, more user-friendly workflow to obtain high transduction efficiency with T cells.
Poster presentations
Smartphone-based titration of AAV vector preparations
With over 130 clinical trials and eight approved gene therapy products, AAV has emerged as one of the most reliable and versatile tools for delivering therapeutic DNA directly into living organisms. The AAV expression system offers several advantages, including efficient transduction, broad tropism, stable high yields, and compatibility with gene-editing components. Quantitative real-time polymerase chain reaction (ddPCR or qPCR), ELISA, cell-based infectious unit assays, and transmission electron microscopy (TEM) have all been used to determine AAV titers. However, these methods are time-consuming and labor-intensive. In this work, Tom Quinn presented an iOS and Android-compatible smartphone application that analyzes a serotype-specific lateral flow assay and delivers particle number values in 10 minutes. The simplicity of the assay facilitates easy monitoring and optimization of AAV production processes to ensure consistency and confidence in downstream applications.
Simple CAR-T cell production: RetroNectin can promote simultaneous T-cell activation and transduction in a G-Rex Bioreactor
Expedited quantification of AAV titers using single-wash ELISA assay

Adeno-associated viruses (AAVs) serve as essential vectors in gene therapy and biomedical research, requiring precise and efficient titer quantification methods to ensure experimental consistency and therapeutic success. In this poster, Yi Zhao presented a single-wash, automation-friendly Enzyme-Linked Immunosorbent Assay (ELISA) protocol for rapid, reproducible quantitation of AAV titers for serotypes 2, 8, and 9. The single-wash approach minimizes hands-on time and procedural complexity, taking only 90 minutes to complete, which is significantly less time than conventional ELISA methods. Validation studies demonstrated a high degree of concordance with quantitative PCR (qPCR) titer data, underscoring its reliability. Its optimized protocol and validated accuracy make it a valuable tool for advancing AAV-based gene therapy research and production, addressing the field's growing demand for efficient and reliable methodologies.
Streamlined ex vivo engineering of human T cells: A single-step approach to T-cell activation and lentiviral transduction
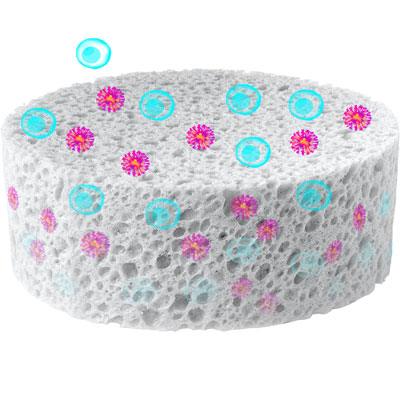
Efficient T-cell engineering is crucial for the success of CAR T-cell therapy but requires multiple labor-intensive steps, including T-cell isolation, activation, and transduction. Herein, we presented a streamlined approach to enhance the ex-vivo engineering of human T cells by integrating cell activation and lentiviral transduction into a single, efficient step, reducing time and resource requirements while maintaining high performance—the Lenti-X T-Cell Transduction Sponge. Our approach leverages a lyophilized alginate-based sponge that is embedded with activation reagents, including anti-CD3 and anti-CD28 antibodies, as well as interleukin-2. The hygroscopic nature and 20–300 nm pore size of the sponge spatially constrain cells and virus to a small area, facilitating efficient transduction and ensuring uniform exposure to activation signals, eliminating the need for spinoculation. The release of activated and transduced cells is facilitated by a chelating release buffer that depolymerizes the sponge, ensuring the rapid recovery of high-quality transduced cells with minimal handling. Our data across multiple donors and operators consistently indicate strong activation and transduction efficiencies, with no adverse effects on growth rates, cell phenotypes, or exhaustion marker expression compared to standard protocols. By providing a streamlined and user-friendly system for T-cell engineering, we strive to drive progress in T-cell therapies and expand their availability to a wider range of patients.
Efficient neuronal transduction of AAV2-derived CereAAV.YN Vector in cynomolgus macaque brain without liver transduction by systemic injection
Featured technologies
Lenti-X Transduction Sponge
The Lenti-X Transduction Sponge facilitates rapid and efficient lentiviral transduction without spinoculation or the use of a chemical transduction enhancer.
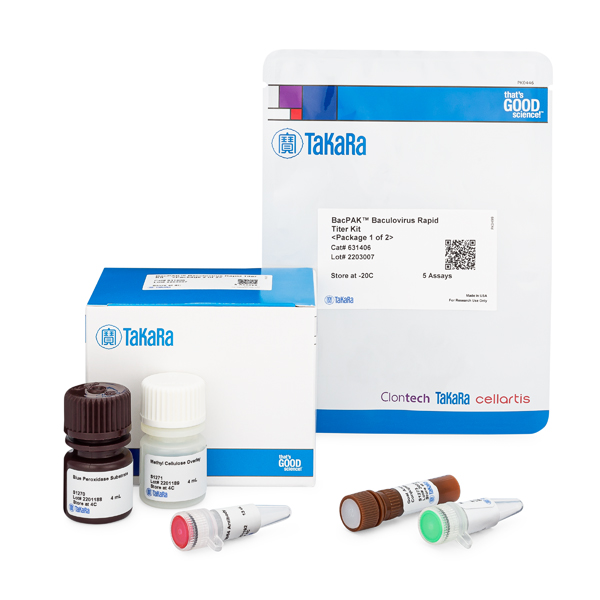
Baculovirus titration kits
Determine baculovirus titers and viral genome content with BacPAK titration kits.
Nucleic acid purification
Complete solutions for purifying nucleic acids from many sources.
qPCR titration
AAV titration kit that includes reagents for extraction of AAV particles from virus-producing cells and qPCR analysis to deterimine titer.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.