Taking single-cell RNA-seq by STORM: Van Andel Institute scientists answer your questions
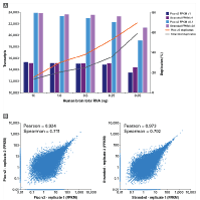
Automated miniaturization of experiments saves on both cost and labor by decreasing reaction volume and minimizing the hands-on work. To this end, research and core facilities at the Van Andel Institute developed Single-cell TOtal RNA Miniaturized sequencing (STORM-seq) as a collaborative effort with Takara Bio. Leveraging the SPT Labtech mosquito HV liquid handler, STORM-seq is a nanoliter-scale, plate-based, single-cell, total RNA approach to yield high-resolution libraries that rival full-volume reactions from both cryopreserved human and mouse tissues.
In this on-demand webinar, Lab Manager Mary Rhodes, Genomics Core Associate II Marc Wegener, and Postdoctoral Fellow Dr. Ben Johnson, describe how the SPT Labtech mosquito HV was used to miniaturize and increase the throughput of the Takara Bio SMART-Seq Stranded Kit to yield robust, total RNA libraries from flow-sorted single cells.
Continue reading for insights from the live Q&A session or register to watch the on-demand webinar here.
Questions and answers
How did you confirm that the SPT Labtech mosquito HV liquid handler produced accurate pipetting volumes?
It took a lot of time and effort. We performed a lot of hands-on testing with water first, then we incorporated different reagents from the kit, specifically with the varying viscosities. We had to confirm the delivery volumes were consistent and within the range we expected. Only then did we finally add samples into the protocol.
How many samples will the Takara Bio SMART-Seq Stranded Kit process with these adaptations for a 384-well plate?
With the SPT Labtech mosquito HV, we have accomplished a six-fold reduction in the original volumes. Ideally, you should be able to do six-fold more reactions than the original kit.
If you are using a 384-well plate later, why are you pooling samples for ribosomal depletion into batches of 96 instead of 384?
We are currently limited by the number of unique indexes we can add to the plates for our pools. As soon as we have access to a larger number of indexes, we will work on creating larger pools of libraries and getting this in even faster throughput.
When considering drop-out rates, does FACS lead to a significant percentage of empty wells in 384-well plates?
When looking at drop-out rates, there are a few things we need to take into consideration.
First, is the potential that the sorter may not successfully place a single cell in each well. From testing at our flow cytometry core facility, we expect less than 10 empty wells in each 384-well plate.
Another thing to consider is negative controls. In our 384-well plates, we include three negative controls that are leading to, in part, that 1% drop-out rate.
Lastly, there could be a potential issue with the genomics portion of the library prep. Altogether, we believe a 1% drop-out rate for the total workflow is very reasonable.
When would you choose STORM-seq over 10x Genomics for single-cell sequencing technology?
We use STORM-seq and 10x Genomics as complementary approaches for answering very different kinds of questions. We use 10x Genomics when interrogating thousands to tens of thousands of cell types to understand the cell-type diversity of our populations.
On the other hand, we use STORM-seq as a targeted, hypothesis-driven approach. You get thousands more genes with STORM-seq compared to 10x Genomics. That can be advantageous to capture some of the intermediate cell types that are otherwise missed with 10x Genomics, especially if these "marker genes" are driven by lincRNAs. A great example of this is the fallopian tube epithelium.
Have you looked at repeat space since you're looking at total RNA (e.g., ERVs, etc.)?
We are in the process of looking at repeat space because we have 2 x 100 reads and are probing the total transcriptome. We don't have data to share yet, but STORM-seq could prove useful in looking at this information.
Does compartmap only work with STORM-seq?
Compartmap also works with single-cell RNA-seq, single-cell ATAC-seq, and whole genome bisulfite sequencing of single cells. It also works with methylation arrays. Beyond that, there are a variety of different single-cell profiling approaches that compartmap is compatible with to provide higher-order chromic confirmation.
What's next for STORM-seq?
In collaboration with Takara Bio and SPT Labtech, we are adding unique molecular identifiers (UMIs) into the mix, as well as some spike-in controls. What originally sparked the development of STORM-seq was the desire to profile the entire transcriptome combined with a multi-omic profile through orthogonal measurements using ATAC-seq or bisulfite sequencing of a single cell. We're working on this all-in-one method, named SCRAP-seq, and it will probably come out next year.
Learn more about our single-cell RNA-seq technologies:
Tech note: stranded libraries from single cells
High-quality, full-length RNA-seq libraries from single-cells with strand-of-origin information.
Tech note: highest sensitivity for single-cell mRNA-seq
View data on the SMART-Seq Single Cell Kit's superior sensitivity and reproducibility for single-cell and nuclei applications.
Webinar: automating full-length single-cell RNA-seq libraries
Register to learn about important considerations for optimizing the automation of single-cell NGS on the Beckman Coulter Biomek.
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.