Gene editing for cancer therapy/drug discovery

CRISPR/Cas9 gene editing has become a powerful method to edit the genomes of many different organisms. First discovered in bacteria as part of an adaptive immune system, CRISPR/Cas9 and modified versions thereof are now broadly used to engineer genomes and to activate or repress the expression of specific genes. Furthermore, CRISPR/Cas9 gene editing promises to accelerate cancer research by providing an efficient technology to dissect mechanisms of tumorigenesis, identify targets for drug development, and possibly arm cells for cell-based therapies (Moses et al. 2018).
Highlighted products
Cancer therapy
Genome editing approaches have enormous potential for targeted, locus-specific cancer treatments. The human papillomavirus (HPV) genes E6 and E7 contribute to the hallmark of resisting cell death by disrupting regular cell cycle and tumor suppressor function. Cas9-mediated HPV E7 oncogene disruption leads to significant inhibition of HPV-induced cancerous activity both in vitro and in vivo, as described by Lao et al., 2018. The authors used our Guide-it Mutation Detection Kit and Guide-it Indel Identification Kit to check for efficient gene editing.

Genome editing approaches have also shown promising results in cancer immunotherapy, to oppose the cancer hallmark of evading immune destruction. Modified chimeric antigen receptor (CAR) T cells have been generated for improved cancer targeting and destruction. Knockin genome modifications in T cells have also been generated with Cas9-sgRNA ribonucleoprotein (RNP) complexes. Kagoya et al., 2018, report that inhibiting DOT1L, a histone H3-lysine 79 methyltransferase, alleviates allogeneic T-cell responses. The authors used the Guide-it sgRNA In Vitro Transcription Kit and Guide-it Recombinant Cas9 (Electroporation-Ready) for CRISPR-mediated TCR ablation in CAR-T cells. Using electroporation of RNP complexes, they could achieve ~30% TCR knockout efficiency in CAR-T cells.
In a groundbreaking study recently published in the journal Nature, Prof. Alexander Marson and colleagues describe a highly efficient method for T-cell engineering that circumvents viral delivery and minimizes cellular toxicity by employing electroporation of Cas9-sgRNA RNP complexes in tandem with double-stranded DNA or single-stranded DNA (ssDNA) HDR templates (Roth et al. 2018).
Drug discovery
Genome-wide knockout screens are a powerful functional genomics tool to discover novel drug targets for cancer therapy. For pooled knockout screens with CRISPR/Cas9, a cell population with a diversity of gene knockouts needs to be generated. Lentiviral particles encoding an sgRNA library are used to infect Cas9-expressing cells at a low multiplicity of infection so that every cell potentially carries a distinct sgRNA cassette and specific gene knockout. Subsequently, this pool of knockout cells is exposed to selected perturbations, followed by NGS analysis compared to a reference control cell population. By this means, it is possible to monitor the phenotypic effect of specific gene knockouts within the cell population.
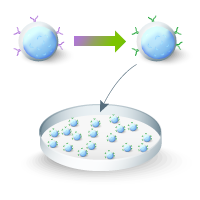
The Guide-it CRISPR Genome-Wide sgRNA Library System is a pooled lentiviral sgRNA library targeting the whole human genome for knockout screens and thus serves as an ideal tool to discover novel drug targets for cancer therapy (Figure 1). The library contains sgRNAs from the Brunello library, based on a recent algorithm for optimized guide sequences for each gene (Doench et al. 2016, Doench et al. 2018):
• four guides per gene
• 76,610 guides in total (includes 172 negative controls)
• 19,114 genes targeted

Figure 1. Schematic of a pooled sgRNA library screen for 6-thioguanine resistance.
References and product citations
Doench J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Doench J. G. et al. Am I ready for CRISPR? A user's guide to genetic screens. Nat. Rev. Genet. 19, 67–80 (2018).
Kagoya Y. et al. DOT1L inhibition attenuates graft-versus-host disease by allogeneic T cells in adoptive immunotherapy models. Nat. Communications 9, 1915 (2018).
Lao Y. H. et al. HPV Oncogene Manipulation Using Nonvirally Delivered CRISPR/Cas9 or Natronobacterium gregoryi Argonaute. Adv. Sci. 1700540 (2018).
Moses C. et al. Hallmarks of cancer: The CRISPR generation. Eur. J. Cancer 93, 10–18 (2018).
Roth, T. L. et al. Reprogramming human T cell function and specificity with non-viral genome targeting. Nat. Lett. 559, 405–409 (2018).
Featured products
Takara Bio USA, Inc.
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.565.6999
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES. © 2025 Takara Bio Inc. All Rights Reserved. All trademarks are the property of Takara Bio Inc. or its affiliate(s) in the U.S. and/or other countries or their respective owners. Certain trademarks may not be registered in all jurisdictions. Additional product, intellectual property, and restricted use information is available at takarabio.com.